Polarized light microscopy is a technology involving polarized light for visualizing biomolecules aligned regularly. The fluorescence polarization microscopy, however, has not been widely used so far. Through the clever application of the newly developed fluorescent labeling technique, it could become a powerful tool for visualizing orientational dynamics of biomolecules, hard-to-reach data with conventional fluorescence microscopies.
The research group led by Professor Sumio Terada, Assistant Professor Keisuke Sato and graduate students including Ayana Sugizaki, Graduate School of Medical and Dental Sciences, Tokyo Medical and Dental University, in a collaborative study with Ochanomizu University, the RIKEN Center for Biosystems Dynamics Research and the Woods Hole Marine Biological Laboratories in the United States, have developed a new fluorescent labeling technique, POLArIS, enabling fluorescence imaging using fluorescence polarization in the life sciences. They first applied it to fertilized starfish eggs, where they found actin filaments radially extending from centrosomes in association with microtubule asters during mitosis, a previously unknown phenomenon. Professor Terada explained that, "Our technique has a potential for fairly broad applications. In particular, when observing a complex biological molecular assembly in a cell, this technique allows us not just to see that it exists but to monitor its dynamics in detail. We are going to explore this possibility further in future." The finding was published in PNAS.
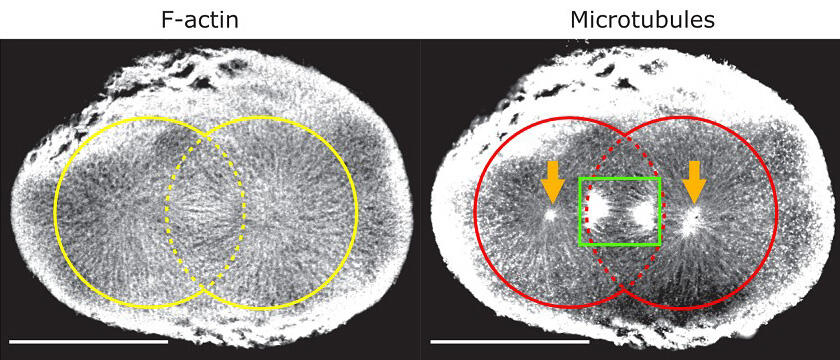
Scale bar, 100 μm. (Credit: Tokyo Medical and Dental University)
Polarized light microscopy, developed by Dr. Shinya Inoue (later a Distinguished Scientist at Marine Biological Laboratories) in the aftermath of World War II at Misaki Marine Biological Station, the University of Tokyo, revealed the vivid, real-time dynamics of mitotic spindles during mitosis within the fertilized eggs of sea urchins - something not visible using the optical microscopy at the time. This observation is recognized as one of the historic achievements in modern cell biology.
Fluorescence microscopes are widely used in today's cutting-edge research fields of life sciences. They have enabled detailed observation of the dynamics of fluorescently labeled biomolecules. While current fluorescence microscopies use fluorescence properties such as intensity, spectrum and lifetime, the information gained from fluorescence polarization is rarely employed. To make use of fluorescence polarization, the target molecule needs to be labeled with a fluorescent molecule through a "rigid" connection, but due to the difficulties in predicting the 3D- structure of a fusion protein, such labeling has been achieved only in special cases.
In order to solve this problem, the research group used Affimer, an artificial antibody-like small molecule that can be expressed intracellularly. Affimer is small and highly stable, and can bind to a variety of targets with high affinity. If Affimer can be "rigidly" connected to a fluorescent protein, then it can be used as a scaffold for the fluorescent protein to be "rigidly" attached to the target molecule of interest.
The amino-terminus of Affimer has an α-helix (secondary structure of protein), while the green fluorescent protein (GFP) amino-terminus has a 3.10 helix (secondary structure of protein). By creating a GFP variant called a circular permutant, the research group produced a GFP with the 3.10 helix at the carboxyl-terminus, the opposite end to the amino-terminus. The fusion protein created by connecting the helices at the carboxyl-terminus of the GFP variant and the amino-terminus of the Affimer was designated as POLArIS (Probe for Orientation and Localization Assessment, recognizing specific Intracellular Structures of interest).
The team first prepared POLArIS that specifically binds to polymerized actin, one of the cytoskeletal proteins, and confirmed that the orientation of fluorescence polarization of the probe indeed reflected the orientation of the actin polymer. POLArIS can in principle be used as a fluorescent polarization probe for any target biomolecules of interest. Sato says, "We were able to observe the orientational dynamics of actin filaments at the single-molecular level in living cells."
Furthermore, when the POLArIS for actin polymer was expressed in a fertilized starfish egg and the following actin dynamics were observed through a fluorescence polarization microscope, the radially extended actin architecture was visualized, that repeatedly appeared and disappeared during every cell division.
This actin structure extended from centrosomes in association with radially extending microtubules, which are known to appear in mitosis, with some associated with mitotic spindles. Actin filaments running along the microtubules of mitotic spindles have been reported recently, but it has not been documented that dynamic actin filaments also extend along astral microtubules, growing out from centrosomes to cell surface.
The research group designated the actin architecture as the FLARE (FLuffy And Radial actin-aster associated with mitosis in Embryo) structure. The dynamics of microtubules during mitosis have been extensively studied and are described even in high school biology textbooks. However, the FLARE structure accompanying the microtubule structure during mitosis was not known. The functional significance of the FLARE structure remains to be determined.
Since POLArIS is genetically encoded, it can be used to observe a wide range of living specimens, from cultured cells to developing embryos and animals. It can also be applied to various observation techniques including microscopy and is expected to promote the development of new fluorescence spectroscopic devices.
This article has been translated by JST with permission from The Science News Ltd.(https://sci-news.co.jp/). Unauthorized reproduction of the article and photographs is prohibited.




