The pathological conditions involving immune cells are diverse, and include immune disorders, infectious diseases, cancer, and arteriosclerosis. Many gene polymorphisms involved in disease onset have been identified; however, understanding how polymorphisms are involved in disease onset requires a large-scale database combining cellular gene expression and genomic sequencing.
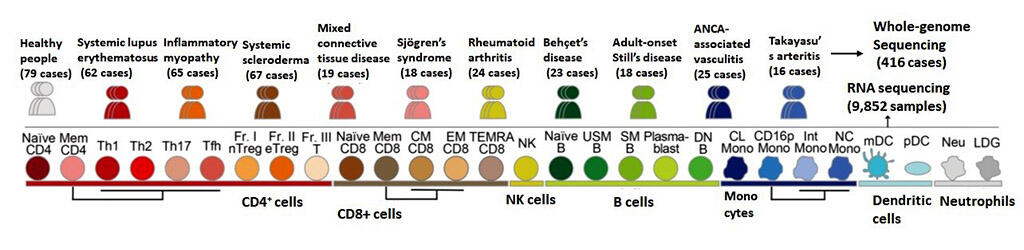
A research team led by specially appointed Assistant Professor Mineto Ota and Professor Keishi Fujio of the Graduate School of Medicine at the University of Tokyo analyzed 9,852 immune cell samples comprising 28 types collected from the peripheral blood of 416 patients to build a gene expression database much larger than those reported in past studies. They then analyzed its relationship with the genome sequences, creating a catalog of functions of gene polymorphisms. Combined with previously reported GWAS data (genome-wide association studies), they revealed cell types and genes associated with various immune disorders.
The database of genome functions has been built mainly with European and American patients; however, the Japanese catalog created in this study is attracting attention for developing genomic research targeting Asians, as well as for detailed understanding of genomic functions through integrated analysis with data on the European and American populations in the future.