In collaboration with the National Institute of Genetics, the research group of Satoshi Hiraoka, a researcher of the Research Center for Bioscience and Nanoscience (CeBN), Research Institute for Marine Resources Utilization, Japan Agency for Marine-Earth Science and Technology (JAMSTEC), analyzed seawater samples collected off the coast of the Boso Peninsula and succeeded in clarifying extensively the DNA chemical modification of the genome (epigenome) of marine microbial flora.
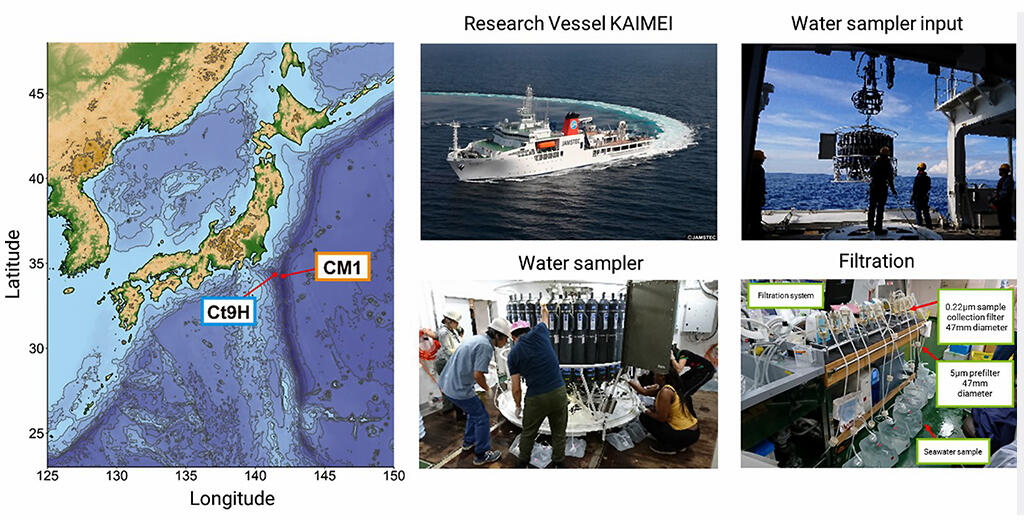
Provided by JAMSTEC with modifications
In bacteria, archaea, and double-stranded DNA viruses, genomic DNA is subjected to various chemical modifications (epigenomes) as in eukaryotes such as humans. However, the diversity of microbial epigenomes and their roles are mostly shrouded by mystery, because most of the microbes in the ocean and other environments are difficult to analyze as they cannot be cultivated in the laboratory. In September 2019, the research group used the Research vessel "KAIMEI" operated by JAMSTEC to collect a large amount of water from the sea area 50 kilometers off the Boso Peninsula. Four layers of microbial samples, ranging from 5 meters on the surface to 300 meters deep, were collected via filtration.
Shotgun sequencing (a sequencing procedure applied to the sequencing of long DNA) was performed on these samples using three types of sequencers, and sequence data was acquired. Through various bioinformatics analyses, they acquired draft genomes (P-MAG) of 233 prokaryotes and draft genomes (V-MAG) of 163 double-stranded DNA viruses. Using PacBio (sequencer) reads, they detected DNA chemical modifications in the genome, predicted their modification motifs, and found a total of 220 diverse modification motifs, including many novel ones. A variety of patterns were observed from prokaryotic genomes, including those with epigenomes that varied widely even between systematically close genomes, those with highly similar epigenomes even within relatively distant lineages, and those with no detectable DNA chemical modifications. In addition, in the viral genome, the lineage group in which DNA chemical modification was observed (Myoviridae, etc.) and the lineage group in which DNA chemical modification was not observed were clearly separated.
Thereafter, the analysis focused on a DNA chemical modification called DNA methylation. A total of 276 DNA methylation enzyme (MTase) genes were predicted as enzymes that undergo this type of chemical modification. A total of 11 MTase enzymes, including enzymes that specifically recognize five novel motifs, were identified from artificial gene synthesis, recombination experiments with Escherichia coli, and experiments with isolated and purified enzymes. In addition, they focused on Alphaproteobacteria, the most abundant microbial lineage in the ocean for more detailed genomic analysis and found that the recognition motif of DNA methylase changed during evolution in the lineage group. Moreover, it was found that the base appearance pattern of the entire genome changed according to the change. This suggests that the epigenome and genome have coevolved.
The researchers have shown that DNA chemical modifications, despite being like an ornamental accessory attached to the genome, can be one of the important factors that greatly affect the genome itself and drive microbial evolution. Dr. Hiraoka said, "In the future, we will clarify the diversity of microbial epigenome on a global scale by analyzing a wide range of microbial flora from places such as deep-sea sediments and terrestrial environment, in addition to those obtained from seawater. I want to comprehensively clarify the impacts of these factors on evolution and ecology."
This article has been translated by JST with permission from The Science News Ltd.(https://sci-news.co.jp/). Unauthorized reproduction of the article and photographs is prohibited.




