A research team led by Professor Hidemasa Nakaminami of the Department of Clinical Microbiology, School of Pharmacy, Tokyo University of Pharmacy and Life Sciences (TUPLS), and Associate Professor Takeaki Wajima of the Faculty of Pharmacy, Meijo University, in collaboration with Professor Yoshiaki Kawamura of the Laboratory of Microbiology, School of Pharmacy, Aichi Gakuin University, has discovered a new species of streptococcus that is resistant (multi-drug resistant) to a variety of antibacterial drugs. The bacterium was named Streptococcus toyakuensis after the Internet domain name of Tokyo University of Pharmacy and Life Sciences, "toyaku." Streptococcus is a known endemic human bacterium but because it is isolated from the blood, it can be highly pathogenic. Furthermore, it was revealed that the drug resistance of this bacterium is transmitted horizontally to closely related Streptococcus spp., so it is necessary to pay attention to future outbreaks. The team's findings were published in the Journal of Global Antimicrobial Resistance.
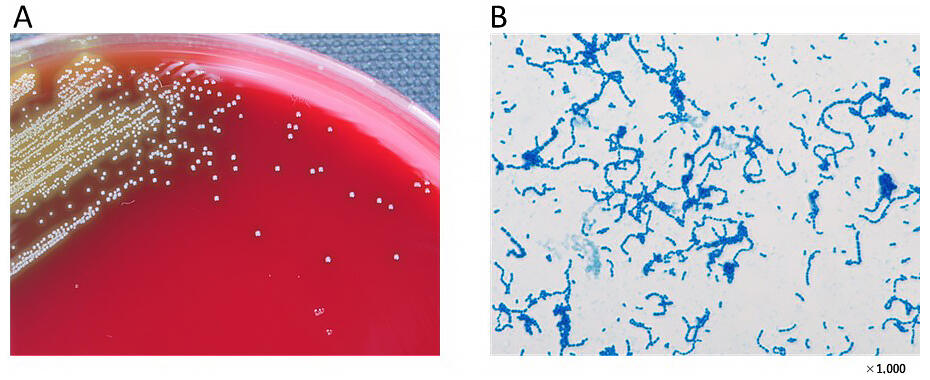
Provided by TUPLS
In recent years, the emergence of drug-resistant bacteria that are resistant to antimicrobial agents has become a problem. Drug-resistant bacteria are emerging not only in pathogenic bacteria but also in commensal bacteria. Streptococcus oralis, a well-known example of a human commensal bacterium, belongs to the genus Streptococcus and is also known as α-haemolytic streptococci or viridans streptococci. Most α-haemolytic streptococci are indigenous in the oral cavity, such as Streptococcus mitis and S. oralis, but there are also bacteria that cause respiratory tract infections and otitis media, such as Streptococcus pneumoniae.
The research group isolated the α-haemolytic streptococci strain TP1632, belonging to the multidrug-resistant mitis group, from the blood of hospitalized patients in 2018. Since the species could not be identified by conventional methods, they conducted genomic analysis, biochemical characterization tests and phylogenetic analysis to analyze the species.
Homology analysis of 16SrRNA gene sequences and comparative genomic analysis showed that it is a new species that is most closely related to S. mitis and at the same time different from existing fungal species by international standards. Biochemical characterization studies also revealed that the use of D-Lactose, D-Raffinose and N-acetyl-β-glucosaminidase was different compared to S. mitis. Since the bacterium was isolated and identified at the Tokyo University of Pharmacy and Life Sciences, it was named Streptococcus toyakuensis after "toyaku," the Internet domain name of the university.
Multidrug-resistant α-haemolytic streptococci classified in the mitis group, such as this bacterium, are known to act as acceptors or donors of drug resistant genes, regardless of their virulence, and contribute to the resistance of other pathogenic bacteria. In fact, drug resistance of the bacteria was also found to be transmitted to other mitis groups. The genome sequence data and strains are stored in public databases and microorganism repositories accessible to researchers worldwide.
This article has been translated by JST with permission from The Science News Ltd.(https://sci-news.co.jp/). Unauthorized reproduction of the article and photographs is prohibited.




