The joint research group of Research Fellow Hiro Nakamura, Senior Researcher Tamao Hisano, and Team Leader Mikako Shirouzu of the Laboratory for Protein Functional and Structural Biology, RIKEN Center for Biosystems Dynamics Research, and Specially Appointed Professor Yoshitsugu Shiro and graduate student Md. Mahfuzur Rahman of the Graduate School of Science, University of Hyogo, have elucidated the mechanism of the "ABC (ATP-binding cassette) transporter, HrtBA" that expels host-derived heme, which is toxic to pathogenic bacteria (Gram-positive bacteria).
Courtesy of RIKEN
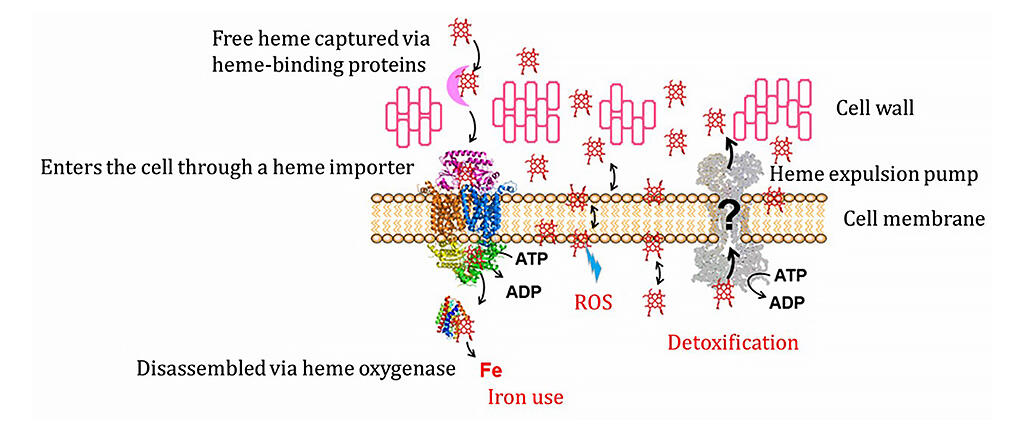
Iron is an essential element for most living organisms. Most of the iron in living organisms exists as heme (iron protoporphyrin IX), in which a ring-shaped organic molecule (porphyrin) covers iron ions. By binding to various proteins, heme is responsible for many of the biochemical reactions that are essential for life activities, such as oxidation-reduction reactions and oxygen transport. On the other hand, it is also known that free heme not bound to proteins has cytotoxicity because it generates reactive oxygen species.
The research group verified whether the HrtBA protein possessed by Staphylococcus aureus, a Gram-positive bacterium, has the ability to detoxify heme. The K12 laboratory strain of E. coli bacteria is resistant to heme because it is encased in an outer membrane that is impermeable to heme. When recombinant DNA technology was used to introduce external heme into the cells of this E. coli strain, their growth rate was reduced, confirming that heme is toxic to the strain. Further expression of the HrtBA gene from diphtheria bacteria in this heme-sensitive recombinant strain restored its growth rate.
Next, in order to investigate the properties of the HrtBA protein as a membrane protein, it was solubilized and purified from the recombinant E. coli bacteria cell membranes in the presence of a surfactant and was embedded back into lipid nanodisks to create a psuedo-cell membrane in which only the HrtBA protein was embedded. The research group then confirmed that the addition of heme to the HrtBA protein that was reconstituted into nanodisks promoted ATP hydrolysis activity of the HrtBA proteins. Additionally, it was also found that heme was released from the HrtBA proteins when ATP was added to HrtBA proteins bound to heme.
The results showed that the ABC transporter is an efflux pump that traps free heme that has entered the cell membrane of pathogenic bacteria and then pushes the heme out through a conformational change that is caused by its subsequent binding to ATP. This experimental data suggests that the heme expelled by the HrtBA protein proceeds via the following three reactions, being pumped out of the cell membrane rather than being transmitted through the cytoplasm to the cell membrane.
(1) When HrtBA proteins are not bound to ATP, they bind to heme that has entered the cell membrane and extract the heme from the membrane.
(2) When ATP binds to HrtBA proteins bound to heme, the bound heme is expelled via structural conversion of the transmembrane domain of the HrtBA protein.
(3) The ATP is degraded by the ATP hydrolysis activity of the HrtBA protein.
Thus, the research group derived a model in which the HrtBA protein structure reverts to (1) when it reaches state (3), and the function of the heme pumps turns over.
Nakamura said that, "As is well known for hospital-acquired infections, even indigenous/resident bacteria such as Pseudomonas aeruginosa (Gram-negative bacteria), Staphylococcus aureus, and Streptococcus (Gram-positive bacteria) can cause life-threatening sepsis and meningitis if they develop multidrug resistance and do not respond to conventional antibiotics, so we hope that this research leads to the development of new antibiotics. Although functional inhibitors targeting HrtBA are limited to Gram-positive bacteria, they are expected to have an antibacterial effect by inhibiting the proliferation of bacteria in the blood."
This article has been translated by JST with permission from The Science News Ltd.(https://sci-news.co.jp/). Unauthorized reproduction of the article and photographs is prohibited.




