A research group led by Professor Norio Ozaki at the Graduate School of Medicine, Nagoya University, Dr. Itaru Kushima and Assistant Professor Miho Imaeda at Nagoya University Hospital, and Deputy Director Satoshi Tanaka at the National Hospital Organization Higashiowari Hospital, announced on August 10 that that genomic copy number variations (CNVs) are involved in the risk of developing eating disorders through genome analysis they carried out related to the disorders. They found risk CNVs known to be associated with neurodevelopmental disorders in 10% of patients with severe eating disorders, many of which affected genes associated with neuronal synapses. Gene set analysis also confirmed that patient CNVs clustered significantly more in the synapse-related gene cluster, demonstrating for the first time that synaptic dysfunction is deeply involved in the pathogenesis of eating disorders. Their findings are expected to lead to the development of treatments for eating disorders. The research was published in Psychiatry and Clinical Neurosciences.
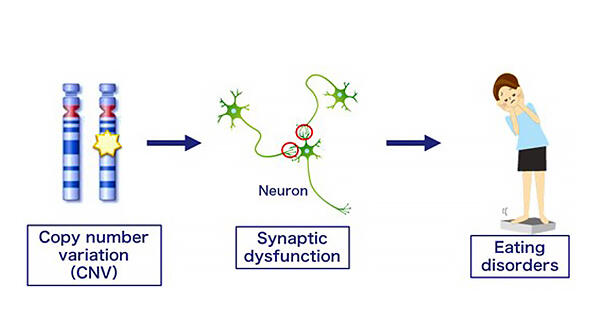
Provided by Nagoya University
Eating disorders are a group of disorders that affect the body and mind due to persistent behavioral abnormalities related to eating, with anorexia nervosa (AN) prevalent among women, affecting about 1% of them. Patients with AN suffer from a distorted body image, resulting in low body weight due to dietary restrictions and other factors. The mortality rate of AN patients is high at about 5% per decade. However, the pathogenesis is unknown, and no effective drug treatment exists. Previous epidemiological studies have shown that genetic factors are strongly involved and may overlap with genetic factors in other psychiatric disorders. A genomic study of eating disorders found CNVs, a subtype of genomic variation associated with the risk of neurodevelopmental disorders such as autism spectrum disorder and psychiatric disorders such as schizophrenia, in a subset of patients, but the association has not been definitive.
As such, the research group analyzed CNV in 70 patients with eating disorders and 1,036 healthy subjects to identify the relationship between eating disorders and CNV. The patients were severely ill (all Japanese women) with a minimum BMI of 15 or less, diagnosed with either AN (restricted eating or overeating and purging) or avoidant/restrictive food intake. A high-resolution array CGH, which can detect even small-sized CNVs, was used to conduct the gene analysis.
A CNV analysis focusing on CNVs involved in neurodevelopmental disorders that had already been identified found the risk of CNVs for neurodevelopmental disorders in 10% (7/70) of patients with eating disorders and 2.3% (24/1036) of healthy subjects. Statistical analysis confirmed that they were significantly associated with the risk of eating disorders (odds ratio = 4.67). Mutations found in patients included Turner syndrome (45, X), KATNAL2 deletion, DIP2A deletion, PTPRT deletion, RBFOX1 deletion, CNTN4 deletion, MACROD2 deletion and FAM92B deletion.
Of these, PTPRT, DIP2A, RBFOX1, and CNTN4 were found to be related to synaptic function. The research group then analyzed their involvement.
The gene set analysis revealed that patient CNVs clustered significantly more (odds ratio = 2.55) in a group of genes related to synaptic signaling. "Eating disorders are a disease for which there are still many unknowns, and it is necessary to identify the pathogenic mechanism in order to develop a treatment for these disorders," explains Dr. Kushima. "We believe that our findings are just the first step. We have been analyzing the genomes associated with psychiatric and developmental disorders and have found common mutations between these disorders and eating disorders. Moving forward, we would like to increase the number of samples, identify more strongly related mutations, and analyze them using animal models in order to develop treatments."
Journal Information
Publication: Psychiatry and Clinical Neurosciences
Title: Contribution of copy number variations to the risk of severe eating disorders
DOI: https://onlinelibrary.wiley.com/doi/10.1111/pcn.13430?af=R
This article has been translated by JST with permission from The Science News Ltd.(https://sci-news.co.jp/). Unauthorized reproduction of the article and photographs is prohibited.




