A research group led by Kohji Yamada, a lecturer from the Department of Biochemistry, Jikei University School of Medicine, and Tsunekazu Oikawa, a lecturer from the Division of Gastroenterology and Hepatology, Department of Internal Medicine, Jikei University School of Medicine, has discovered a novel secretion mechanism by which some cytosolic proteins are released from liver cancer cells to the extracellular matrix via the endoplasmic reticulum, and revealed that a part of these proteins, protein kinase C delta (PKCδ) in serum may be useful as a biomarker for early cancer diagnosis. Comprehensive proteomic analysis revealed that secreted cytoplasmic proteins localize within the endoplasmic reticulum. The research group identified the secretion mechanism to the extracellular space via interaction with endoplasmic reticulum inner membrane molecules. The results are expected to reveal the mechanism of the onset of liver cancer and its early diagnosis and were published in the August 31 issue of PNAS and the August 4 issue of Gastro Hep Advances.
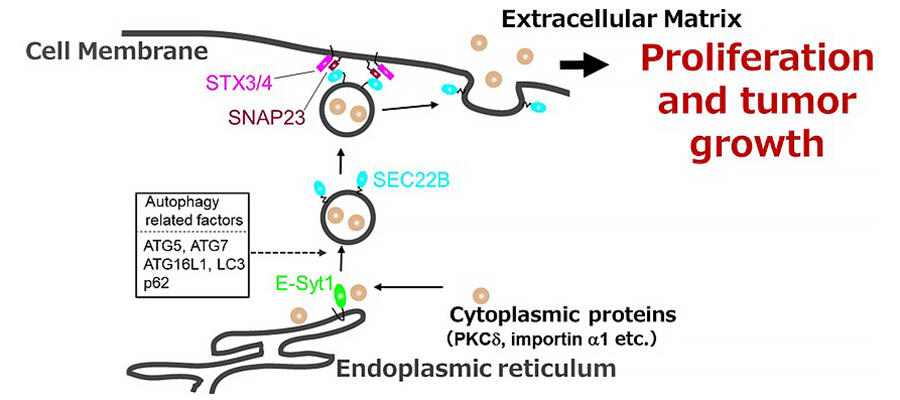
(i) Some cytoplasmic proteins localize to the endoplasmic reticulum by interacting with ER membrane protein E-Syt1.
(ii) Autophagy related factor-dependent uptake into SEC22B-positive vesicles. (iii) Released from the plasma membrane via SNAP23, STX3, and STX4, which are plasma membrane SNAREs.
Provided by the Jikei University School of Medicine
Liver cancer is a disease that affects about 38,000 people per year in Japan and kills 25,000. It is known that subjective symptoms are difficult to detect, and that the prognosis becomes worse as the disease progresses. Liver cancer is known to develop in stages through chronic hepatitis, non-alcoholic steatohepatitis and cirrhosis, but as with other cancers, the mechanism behind the onset of the disease has remained a mystery.
The research group had previously discovered that proteins such as the nuclear transport factor importin1 and PKCδ, which remain intracellular in normal cells, are specifically secreted extracellularly in liver cancer cells, and that these proteins have cell proliferation potential. In this study, the research group focused on PKCδ and performed a comprehensive proteomic analysis to identify the secretion mechanism.
As a result, they found PKCδ localizes to the endoplasmic reticulum by interacting with ER membrane protein E-Syt1. Suppression of E-Syt1 expression reduced this localization, confirming the absence of such localization in gastric cancer cells that do not secrete importin1 or PKCδ. They also found that PKCδ is taken up by autophagic vesicles (SEC22B) through interaction with E-Syt1 and released into the plasma membrane through interaction with SNARE proteins on the plasma membrane.
Furthermore, when an antibody reagent for PKCδ was delivered intracellularly and its effect on tumorigenicity was examined, this antibody suppressed it. This demonstrated the potential for its use as a new therapeutic target.
Based on these basic studies, the research group then investigated the potential use of PKCδ in serum of patients with liver cancer as a blood biomarker for the early diagnosis of liver cancer. First, they measured the serum of nine healthy subjects and 313 clinical patients with chronic liver disease (including 182 cases of various stages of liver cancer) using a sandwich ELISA to verify its diagnostic ability and compare it with two existing liver cancer markers (AFP and PIVKA2).
The results showed that protein kinase PKCδ in the liver cancer group was significantly higher than that in healthy subjects and patients with chronic liver disease. They found that protein kinase PKCδ can be used as a novel biomarker to distinguish patients with liver cancer from healthy controls. The diagnostic ability of protein kinase PKCδ in chronic liver disease was comparable to that of existing markers for liver cancer.
The results indicate that combining the two may improve diagnostic performance since there is no association between protein kinase PKCδ and each of the existing markers. The researchers expect to be able to detect about half of the cases where the two existing markers are negative.
Furthermore, unlike the other two existing markers, protein kinase PKCδ has a higher positivity rate for very early-stage liver cancers smaller than 20 mm (about 45%) than the existing markers (about 15% each), indicating that it may help with early diagnosis, which has been elusive.
The research group plans to develop a diagnostic kit, analyze the relationship between protein kinase PKCδ and patients' clinical and medical characteristics, and verify its usefulness through multicenter collaborative studies.
This article has been translated by JST with permission from The Science News Ltd.(https://sci-news.co.jp/). Unauthorized reproduction of the article and photographs is prohibited.




