The research group of Professor Satoru Konnai and Specially Appointed Assistant Professor Tomohiro Okagawa of the School of Veterinary Medicine, Hokkaido University, Masumichi Saito of the National Institute of Infectious Diseases, Takahiro Matsudaira of FASMAC, and Professor Kenji Murakami and Associate Professor Shinji Yamada of the Faculty of Agriculture, Iwate University, have developed a clonality analysis technique for virus-infected cells and applied it to diagnosing and predicting the onset of bovine infectious lymphoma.
Provided by Hokkaido University
Bovine Lymphoma Virus (BLV) causes lymphoma (EBL) in cattle, and, when EBL is detected, all of the affected cattle are forcibly culled, resulting in a huge loss for the farmer, not just in lost sales revenue when the rest of the cattle are sold, but also in the loss of all of the time and money that they have spent up to that point, including the costs for feed, breeding, and labor, and so on. Currently there is no effective vaccine or treatment for BLV, and even though isolation measures are taken based on current infection diagnoses, in many cases they are not effective. As a result of this, a method of predicting the disease's onset has been sought.
The research group used a comprehensive amplification method (RAISING) for the provirus (a viral genome integrated into the host cell's DNA) insertion site of BLV, and then verified the amplification sensitivity and reproducibility of the method. Using blood samples from 287 BLV-infected cattle from domestic farms (AL asymptomatic stage, PL lymphocyte-enriched stage, EBL lymphoma, and death due to systemic lymphoma) and 169 tumor samples, the research group amplified the provirus insertion site via RAISING, and then used analysis software (CLOVA) to analyze the clonality value (Cv), which is an index showing the degree of cloning of BLV-infected cells.
The results showed that RAISING targeting BLV is highly sensitive and accurate, and that it can measure Cv in infected cells with a high degree of reproducibility. Cv analysis of BLV-infected cattle showed that Cv was higher in EBL than in uninfected cattle (AL and PL). However, provirus levels, a conventional diagnostic marker, were similar in EBL and PL blood samples, and no clear difference was observed. In fact, when EBL diagnosis was attempted using Cv as a marker, EBL could be differentiated with extremely high accuracy, with a sensitivity of 87.1% and a specificity of 93.0%.
On the other hand, when provirus content (number of copies of the provirus) was used as a marker for EBL diagnosis, the sensitivity and specificity were 44.6% and 67.2%, respectively, and it was not possible to distinguish between EBL and unaffected cattle. In addition, samples with high Cv were also identified in some uninfected cows. These cases are currently being followed up on to verify whether or not they developed EBL.
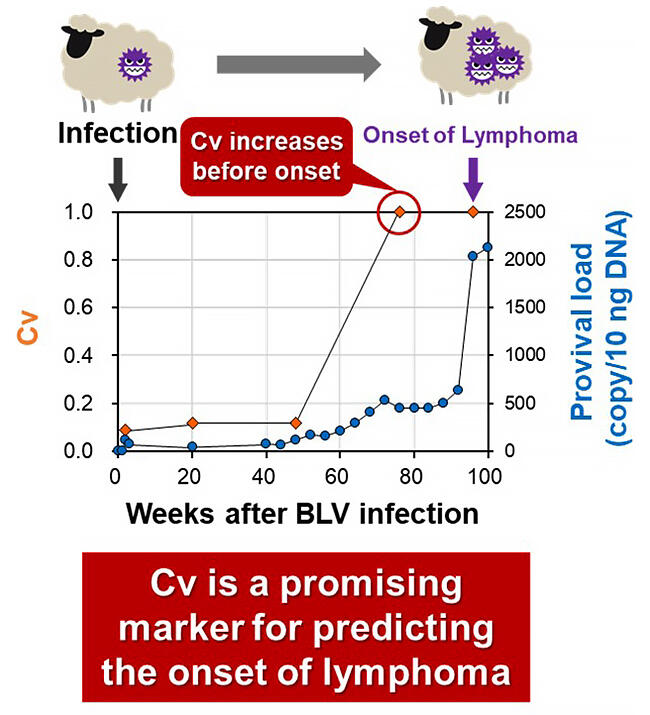
According to Konnai, "Because it takes a long time for cattle to develop the disease after infection, we conducted a longitudinal analysis using sheep, which are an experimental model for EBL infection, and the results showed that this method is useful as a method for predicting disease onset. In the future, we would like to demonstrate the usefulness of the technology in clinical settings through large-scale field studies, and at the same time, we would like to make the analysis kit commercially available so that they can be widely used to reduce the damage caused by this disease in livestock production and improve productivity.
Journal Information
Publication: Microbiology Spectrum
Title: Diagnosis and early prediction of lymphoma using high-throughput clonality analysis of bovine leukemia virus-infected cells
DOI: 10.1128/spectrum.02595-22
This article has been translated by JST with permission from The Science News Ltd.(https://sci-news.co.jp/). Unauthorized reproduction of the article and photographs is prohibited.




