A research group consisting of Professor Masato Katahira, Associate Professor Takashi Nagata and Assistant Professor Yudai Yamaoki of the Institute of Advanced Energy, Kyoto University and their colleagues has announced that they have clarified that the dynamics of nucleic acid base-pairs differ when in vitro. The group created an in-cell Nuclear Magnetic Resonance technique (NMR) that can directly observe nucleic acid NMR signals in living cells and used this to make observations. Through this, they learned that nucleic acid base-pairs open more frequently in living cells than they do in vitro. They also ascertained that non-specific interactions with proteins are the main cause of this opening. It is possible that base-pair opening/ closing is relevant to the effects of nucleic acid medicines, as well as a variety of life phenomena, and it is hoped that this research will further clarify these mechanisms. The group's outcomes were published in the international science journal Nature Communications.
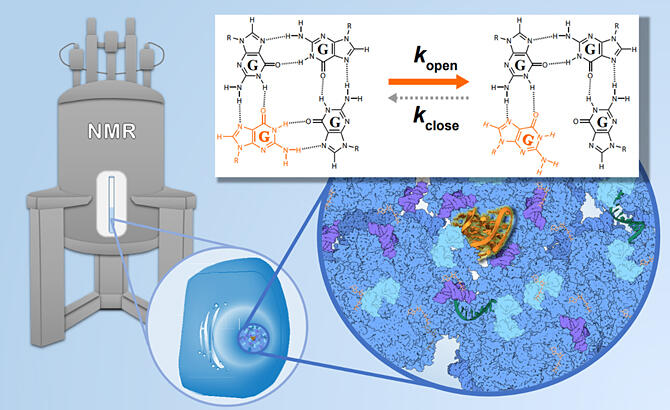
Provided by Kyoto University
Living cells contain 400 grams of proteins and nucleic acid per liter, whereas experiments using test tubes take place in dilute aqueous solution, and it was thought that cell dynamics differ between them. Researchers believed that genetic expression is affected by nucleic acid structure and dynamics such as base-pair opening and closing, but it was difficult to grasp these dynamics due to technical limitations.
In 2018, the research group developed an in-cell NMR technique that enables direct observation of nucleic acid NMR signals in living human cells, thus succeeding in directly observing nucleic acid NMR signals in vivo. Unfortunately, they did not gain an understanding of nucleic acid dynamics in living cells from this.
This time, the research group established a technique that introduced a bio-reactor system to the in-cell NMR, and nucleic acid molecules to living cells. Using the in-cell NMR technique, they were able to take NMR measurements while the cells remained alive for over 24 hours in the NMR equipment, enabling them to obtain large quantities of highly reliable data.
Thus, they used this technology to compare the dynamics of hairpin structure nucleic acid in vitro and in living cells. More specifically, they investigated the speed (frequency) of chemical exchange in vitro and in living cells by making use of the characteristic chemical exchange of the base's imino proton (NH) with the solvent water proton (H), which only occurs when the base-pair is open.
As a result, they learned that nucleic acid base-pairs have a higher opening frequency in living cells than in vitro. They also found that the bases contained in nucleic acid measured with NMR opened with different frequencies. In addition to this, the group investigated quadruplex nucleic acid, which exists in regions key to the regulation of life phenomena (such as promoter regions), and ascertained that, similarly, the opening frequency was higher in living cells.
Next, they explored what the opening frequency rate in living cells is dependent upon. They added glycerol to reduce the amount of water, similar to living cells, and added Ficoll (a macromolecule) to mimic the in-cell density level, but the frequency did not change. They therefore verified the possibility of non-specific interactions resulting from the protein-rich environment within cells. They added negatively charged albumin and positively charged lysozyme respectively.
From this, they discovered that the frequency increased only when lysozyme was added, and that the frequency approached that of living cells depending on the amount added.
These outcomes showed that base-pair opening and closing, which researchers were not previously able to fully clarify, occurs with high frequency in actual living cells, and it is hoped that this will lead to the clarification of the mechanisms of life phenomena associated with it.
Katahira commented, 'We believe that the base-pair opening and closing we have found through this study is an event that triggers change in the spatial structure of nucleic acid through opening. If it becomes possible for us to engage in artificial operations to maintain an open state using low-molecular-weight compounds or nucleic acid that target sites where the base-pairs are open and exposed, I anticipate the possibility that this could lead to the development of nucleic acid medicines and more. In the future, I hope to investigate interactions and spatial structures as well as dynamics - not just those of nucleic acid but those of proteins, too.'
Journal Information
Publication: Nature Communications
Title: Shedding light on the base-pair opening dynamics of nucleic acids in living human cells
DOI: 10.1038/s41467-022-34822-4
This article has been translated by JST with permission from The Science News Ltd. (https://sci-news.co.jp/). Unauthorized reproduction of the article and photographs is prohibited.




