A research group led by Associate Professor Shuji Tani of the Graduate School of Agriculture, Osaka Metropolitan University, has analyzed the regulatory mechanism of carbohydrate hydrolase production in the filamentous fungus Aspergillus aculeatus, which produces enzymes with a high capacity to degrade plant biomass, and discovered that epimerase (Uge5), an enzyme involved in galactose utilization, is involved in the expression control of degrading enzyme genes in filamentous fungi. This has provided some insight into the molecular mechanism by which the expression of enzyme genes in filamentous fungi is selectively regulated in response to the type of sugar that it is induced by.
Provided by Osaka Metropolitan University
The technology to produce various chemical products from the sugars that make up plant cell walls instead of petroleum will greatly contribute to the construction of a sustainable social infrastructure. The research group analyzed the regulation mechanism of carbohydrate hydrolase production in Aspergillus aculeatus, which produces enzymes that are highly capable of degrading plant biomass.
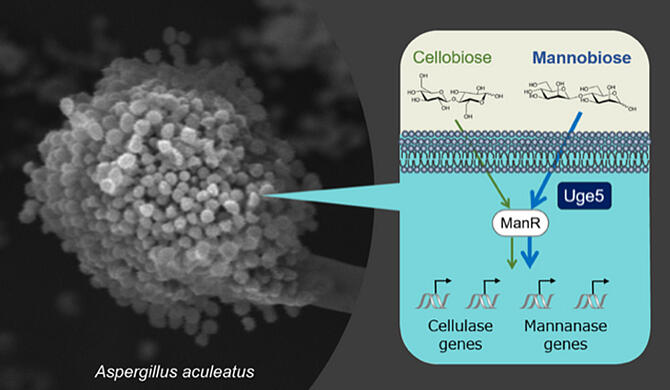
The production of carbohydrate hydrolases in filamentous fungi is regulated by controlling gene expression. In Aspergillus aculeatus, the expression of the plant biomass‐degrading enzymes cellulase and mannanase genes is regulated by the transcription factor ManR. Although high expression of ManR was expected to uniformly increase the expression of various cellulase and mannanase genes, some genes did not increase or decrease in expression, contrary to expectations. The causes of this remain unclear and are a barrier to the development of technologies for the comprehensive mass production of various enzymes.
To identify the mechanism of ManR‐mediated regulation of gene expression, the research group identified one gene involved in ManR‐mediated regulation of gene expression from a library of approximately 9,000 gene disruption strains of this filamentous fungus. The researchers found that gene sequence analysis of the identified genes revealed that epimerase, an enzyme involved in galactose utilization, is involved in the ManR‐mediated gene expression induction in response to mannobiose, but that cellobiose is not involved in gene expression induction. ManR induces cellulase and mannanase gene expression in response to cellobiose and mannobiose, providing an insight into the molecular mechanism by which enzyme gene expression is selectively regulated in response to sugar in filamentous fungi.
"In our research, we discovered that epimerase is involved in the regulation of enzyme production and unraveled a complex, regulated enzyme production mechanism," commented Tani. "We hope to understand the function of epimerase in the future, which will lead to the development of technology for the simultaneous mass production of large quantities of enzymes."
■ Gene disruption strain library: Created by introducing one copy of a DNA fragment of known sequence at any point in the chromosomal DNA of Aspergillus aculeatus. This made it easier to identify disrupted genes.
Journal Information
Publication: Applied Microbiology and Biotechnology
Title: A new function of a putative UDP‐glucose 4‐epimerase on the expression of glycoside hydrolase genes in Aspergillus aculeatus
DOI: 10.1007/s00253-022-12337-8
This article has been translated by JST with permission from The Science News Ltd. (https://sci-news.co.jp/). Unauthorized reproduction of the article and photographs is prohibited.




