A research group led by Professor Takashi Satoh of the Department of Immunology at the Graduate School of Medical and Dental Sciences at Tokyo Medical and Dental University (TMDU), in collaboration with Juntendo University, Osaka University and RIKEN, has revealed the mechanism of severity of novel coronavirus (SARS-Cov-2) infection. Animal experiments and analyses with a large number of patient specimens have shown that patients with severe COVID-19 have high serum concentrations of interleukin-10 (IL-10), which induces alveolar macrophages to express angiotensin-converting enzyme 2 (ACE2), leading to whole lung infection. It was found that the novel receptor 'CiDRE' (COVID-19 infectivity-enhancing dual receptor) is highly expressed in patients with a genetic background of aggravation risk. CiDRE is expected to be used as a biomarker for severity. The findings were published in the July 12, 2023 issue of the international academic journal Immunity.
Provided by TMDU
COVID-19 was first reported in 2019, and the mechanisms of infection and severity still remain unclear.
Alveolar macrophages are immune cells found in the lungs that protect them from foreign invaders such as viruses. Initially, it was thought that the relationship between infection and severity was weak because the novel coronavirus does not induce expression of the ACE2 receptor in cells it infects. However, the novel coronavirus has recently been identified in alveolar macrophages of critically ill patients, and it has been suggested to be involved in the exacerbation of infection.
The research group therefore focused on alveolar macrophages and investigated their involvement in the mechanism of severity.
Hamsters were first infected with the virus and the lungs were observed over time. ACE2 expression confirmed the infection on next day in the upper respiratory tract (nose to throat), but not in the terminal alveoli of lungs; and the infection was finally confirmed in alveolar macrophages on the third day. However, infection was not established in experiments in which the alveolar macrophages isolated from healthy hamsters were directly exposed to the virus.
Since the virus was detectable in the alveolar macrophages over time, the research group examined the possibility that the alveolar macrophages were altered by factors in the surrounding environment which were influenced by the virus infection.
They analyzed clinical samples (blood) obtained from a large number of patients treated at TMDU to investigate cytokines that may alter macrophages. The results showed that IL-10, an anti-inflammatory cytokine, was significantly high at the time of admission in the patients who developed progressive disease after hospitalization. There was no difference in inflammatory cytokines (such as IL-6) between the severe and stable groups.
When alveolar macrophages were cultured with IL-10, they transformed to anti-inflammatory M2c alveolar macrophages, the expression of ACE2 increased rapidly, and the expression of molecules involved in infection and exacerbation was also significantly increased. The viral infection experiments on these macrophages showed that all types of viruses were able to infect them. The research group also confirmed that administration of an IL-10-blocking antibody to infected hamsters significantly inhibited its action and prevented aggravation.
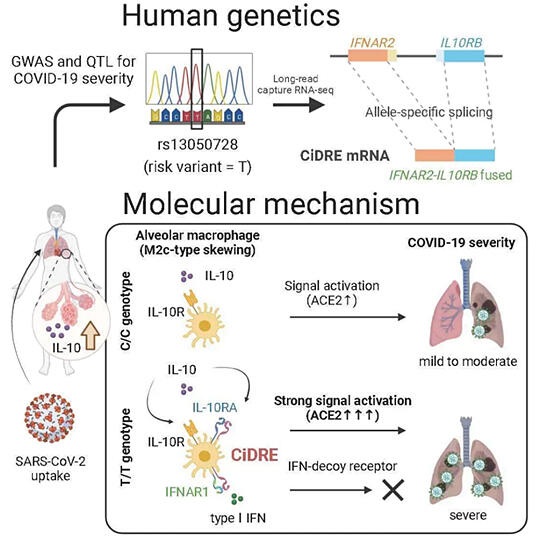
They subsequently performed a genome-wide association study to identify the factors involved in severity and found a single-nucleotide polymorphism (SNP), re13050728, in the IFNAR2 gene, which encodes part of the type I interferon (IFN) receptor, near the IL-10 receptor. Quantitative trait locus analysis revealed that human monocyte-derived macrophages with the T/T genotype of this SNP tended to develop severe disease, whereas those with the C/C genotype did not.
In T/T genotypes, splicing across the two gene regions of IFNAR2 and IL-10 receptors occurs frequently, and the receptor 'CiDRE,' encoded by the newly generated mRNA, is strongly associated with severity of the disease.
By investigating the structure and function of 'CiDRE' in detail, it was found that this receptor binds to IL-10 in alveolar macrophages, enhances ACE receptor expression, promotes viral infection, and simultaneously increases IFN-α. Furthermore, it became clear that aggravation is caused by capturing and attenuating the signal and by nullifying the antiviral response.
IL-10 serum concentration and monocyte CiDRE expression level may be used as biomarkers for aggravation.
Satoh said, "Currently, we are developing a PCR kit to predict infection and severity in a simple and quick manner. The expression level of CiDRE is clearly different between the T/T and C/C genotypes, and this feature is advantageous for determining the risk of aggravation. As there are individual differences in C/T genotypes, we believe that it is possible to make predictions by looking at not only genotypes but also expression levels."
Journal Information
Publication: Immunity
Title: Expression of the readthrough transcript CiDRE in alveolar macrophages boosts SARS-CoV-2 susceptibility and promotes COVID-19 severity
DOI: 10.1016/j.immuni.2023.06.013
This article has been translated by JST with permission from The Science News Ltd. (https://sci-news.co.jp/). Unauthorized reproduction of the article and photographs is prohibited.




