The research group led by Professor Hitoshi Okazawa including Ms. Juliana Bosso Taniguchi (graduate student), Ms. Yuki Yoshioka (graduate student) and Hikari Tanaka (lecturer) (Medical Research Institute, Tokyo Medical and Dental University) in collaboration with Kyoto University, Kyoto Prefectural University of Medicine, and Yokohama City University announced the development of a new treatment concept for Charcot-Marie-Tooth disease (CMT) type 1A (CMT1A), a representative peripheral nerve degenerative disease, using genome editing technology. Using patient-derived induced pluripotent stem cells (iPS cells), they confirmed that it was possible to normalize the duplication of the PMP22 genome region causing CMT1A. The results are expected to lead the development of treatments for this disease and were published in the November 23 issue of the international academic journal Communications Medicine.
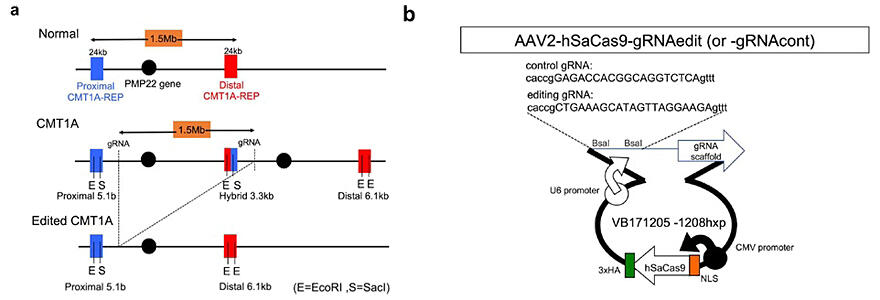
A) Human genome structures surrounding the PMP22 gene of normal, CMT1A, and post-editing CMT1A cells with targets sites for genome editing.
B) Map of the genome editing VB171205 plasmid used to remove additional PMP22 gene copies from CMT1A cells, from which AAV2 vector was generated in HEK293 cells.
Provided by TMDU
CMT is an intractable disease that occurs in approximately 1 out of 10,000 people in Japan. Declined peripheral nerve function causes sensory impairment and muscle weakness of the limbs, which impairs quality of daily life and causes limb deformities in some cases. Approximately 30 causative genes have been identified that comprise various CMT subtypes. Among them, CMT1A accounting for half of the total CMT patients is caused by duplicated long genomic regions (1.5 Mb), containing the causative gene PMP22, on chromosome 17. Owing to the duplication in one chromosome in CMT1A, PMP22 protein expression increases by 1.5-fold and Schwann cells, which require the function of this protein, are thought to become dysfunctional.
Meanwhile, a decrease of this protein is known to cause peripheral neuropathy. Problems arise when the amount of this protein is high or low, making treatment difficult. Previous attempts using nucleic acid drugs have not been successful in clinical trials.
On this background, the research group developed normalization of the CMT1A patient genome by cutting out one of the overlapping genomic regions using genome-editing technology.
First, a guide RNA was designed and then integrated with a genome editing enzyme (Cas9) to create a therapeutic Adeno-associated virus (AAV) vector. Since the region to be excised was long and lacked a verifiable mouse model, they evaluated the genome-editing rate by using iPS cells obtained from CMT1A patients (CMT1A-iPS cells) by Professor Haruhisa Inoue and colleagues (Center for iPS Cell Research Institute and Application, Kyoto University).
When the genome editing rates were validated using both PCR and Southern blotting methods, the rates were between 20% and 40%.
Furthermore, when the patient-derived iPS cells, Schwann progenitor cells, and Schwann cells were examined using the PCR method, the genome editing rates were found to be approximately 40%, 70%, and 40%, respectively, which were greater than expected.
When the phenotypic system was investigated using myelination and cell death as indicators, the results demonstrated that addition of a therapeutic AAV vector to Schwann cells derived from patient-derived iPS cells reduced cell death and restored myelination. Assessment of the PMP22 protein expression level confirmed that addition of AAV vector normalized the expression level.
Examination of off-target mutations further confirmed nearly no off-target mutations in both iPS cells or mice.
The group plans the clinical application of the technique in the future. However, clinical research, including building a financial foundation, may take several years.
Journal Information
Publication: Communications Medicine
Title: AAV-mediated editing of PMP22 rescues Charcot-Marie-Tooth disease type 1A features in patient derived iPS Schwann cells
DOI: 10.1038/s43856-023-00400-y
This article has been translated by JST with permission from The Science News Ltd. (https://sci-news.co.jp/). Unauthorized reproduction of the article and photographs is prohibited.




